Accueil du site > Production scientifique > Comparative Proteomic and Transcriptomic Analysis of Follistatin-Induced Skeletal Muscle Hypertrophy
Comparative Proteomic and Transcriptomic Analysis of Follistatin-Induced Skeletal Muscle Hypertrophy
Date de publication: 15 août 2017
C. Barbé , F. Bray, M. Gueugneau, S. Devassine, P. Lause, C. Tokarski, C. Rolando , J.-P. Thissen
J. Proteome Res. 16 3477–3490 (2017). DOI
Travail réalisé sur le site de l’Université de Lille.
Abstract
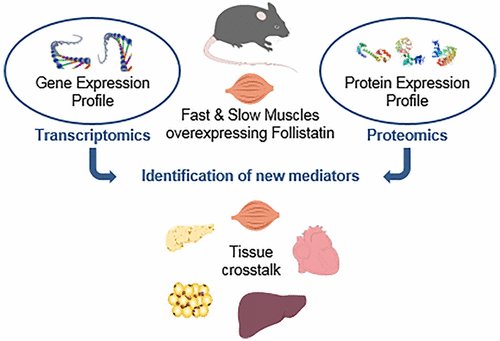
Skeletal muscle, the most abundant body tissue, plays vital roles in locomotion and metabolism. Myostatin is a negative regulator of skeletal muscle mass. In addition to increasing muscle mass, Myostatin inhibition impacts muscle contractility and energy metabolism. To decipher the mechanisms of action of the Myostatin inhibitors, we used proteomic and transcriptomic approaches to investigate the changes induced in skeletal muscles of transgenic mice overexpressing Follistatin, a physiological Myostatin inhibitor. Our proteomic workflow included a fractionation step to identify weakly expressed proteins and a comparison of fast versus slow muscles. Functional annotation of altered proteins supports the phenotypic changes induced by Myostatin inhibition, including modifications in energy metabolism, fiber type, insulin and calcium signaling, as well as membrane repair and regeneration. Less than 10% of the differentially expressed proteins were found to be also regulated at the mRNA level but the Biological Process annotation, and the KEGG pathways analysis of transcriptomic results shows a great concordance with the proteomic data. Thus this study describes the most extensive omics analysis of muscle overexpressing Follistatin, providing molecular-level insights to explain the observed muscle phenotypic changes.








