Accueil du site > Production scientifique > Uncoiling collagen : a multidimensional mass spectrometry study
Uncoiling collagen : a multidimensional mass spectrometry study
Date de publication: 3 novembre 2015
H. J. Simon, M. A. van Agthoven, P. Y. Lam, F. Floris, L. Chiron, M.-A. Delsuc, C. Rolando, M. P. Barrow, P. B. O’Connor
Analyst 141 157-165 (2016). DOI
Travail réalisé sur le site de l’Université de Lille 1, Sciences et Technologies.
Abstract
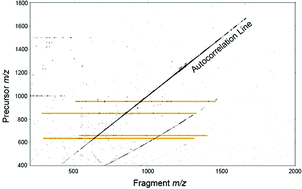
Mass spectrometry can be used to determine structural information about ions by activating precursors and analysing the resulting series of fragments. Two-dimensional Fourier transform ion cyclotron resonance mass spectrometry (2D FT-ICR MS) is a technique that correlates the mass-to-charge (m/z) ratio of fragment and precursor ions in a single spectrum. 2D FT-ICR MS records the fragmentation of all ions in a sample without the need for isolation. To analyse specific precursors, horizontal cross-sections of the spectrum (fragment ion scans) are taken, providing an alternative to conventional tandem mass spectrometry (MS/MS) experiments. In this work, 2D FT-ICR MS has been used to study the tryptic digest of type I collagen, a large protein. Fragment ion scans have been extracted from the 2D FT-ICR MS spectrum for precursor m/z ratios : 951.81, 850.41, 634.34, and 659.34, and 2D FT-ICR MS spectra are compared with a set of 1D MS/MS spectra using different fragmentation methods. The results show that two-dimensional mass spectrometry excells at MS/MS of complex mixtures, simplifying spectra by eliminating contaminant peaks, and aiding the identification of species in the sample. Currently, with desktop computers, 2D FT-ICR MS is limited by data processing power, a limitation which should be alleviated using cluster parallel computing. In order to explore 2D FT-ICR MS for collagen, with reasonable computing time, the resolution in the fragment ion dimension is limited to 256k data points (compared to 4M data points in 1D MS/MS spectra), but the vertical precursor ion dimension has 4096 lines, so the total data set is 1G data points (4 Gbytes). The fragment ion coverage obtained with a blind, unoptimized 2D FT-ICR MS experiment was lower than conventional MS/MS, but MS/MS information is obtained for all ions in the sample regardless of selection and isolation. Finally, although all 2D FT-ICR MS peak assignments were made with the aid of 1D FT-ICR MS data, these results demonstrate the promise of 2D FT-ICR MS as a technique for studying complex protein digest mixtures.








